3D bioprinting has made new headway recently with the publication of a paper titled “Untethered micro-robotic coding of three-dimensional material composition” in Nature Communications in January. Researchers at Brigham and Women’s Hospital and Carnegie Mellon University have teamed up to develop a method for 3D printing biological material using magnetically controlled robots.
The team was made up of BWH’s Dr. Savas Tasoglu, a research fellow in the Division of Renal Medicine, and Dr. Utkan Demirci, associate professor of Medicine in the Division of Biomedical Engineering, along with Drs. Eric Diller and Metin Sitti, professor in the Department of Mechanical Engineering, Carnegie Mellon University. Using what the researchers call “untethered micro-robotic coding”, they were able to remotely control small robots, using magnetic fields, to move hydrogels containing biological material to form specific structures. The magnetic operation of these micro-robots was used so that the construction of objects can be performed without disrupting the vitality and proliferation of biological cells.
To demonstrate the many capabilities of their technique, the researchers performed a number of experiments, such as the arrangement of triangle and rod patterns:
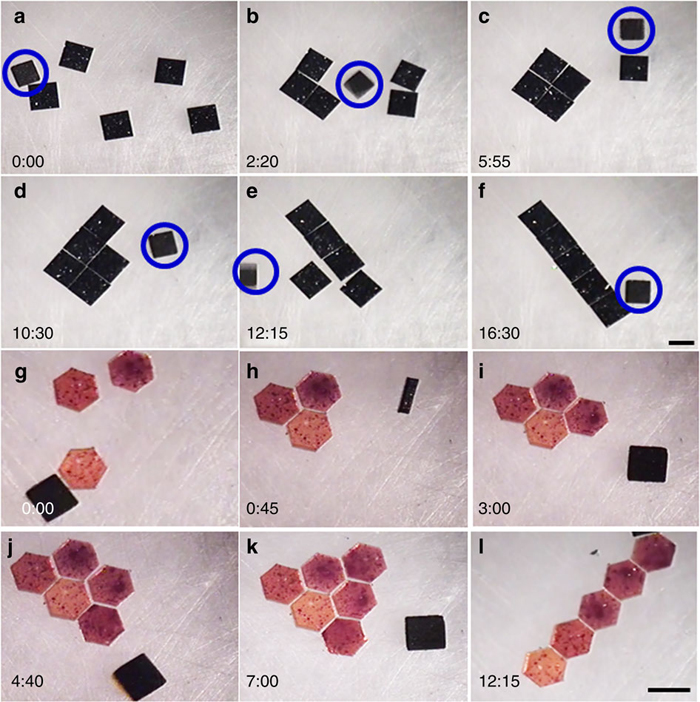
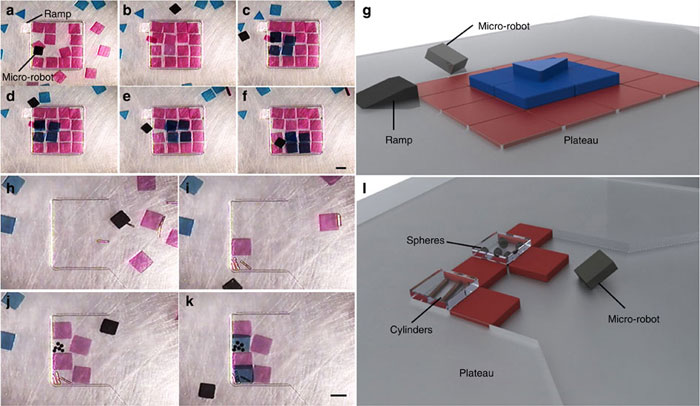
“Micro-robotic coding of (a–f) square silicon chiplets into square and rod patterns, and (g–l) hexagonal polydimethylsiloxane blocks into triangle and rod patterns. All the experiments were performed in a 20 mm × 20 mm × 4 mm chamber. Snapshots of manipulation of 1 mm × 1 mm square silicon chiplets at different time points (shown at the left corner). The time stamp format is minutes:seconds. The magnetic micro-robots are shown in a blue circle (a–f). Black object in each image (g–l) is top-view of the crawling magnetic micro-robot. Scale bar, 1 mm.
And the construction of a pyramid structure made up of differently shaped hydrogel structures and microcomponents, like 100 μm diameter copper cylinders and 200 μm diameter polystyrene spheres:
Micro-robotic creation of (a–g) a three-layer heterogeneous pyramid structure consisting of 16, 4 and 1 gel on each layer, and (h–l) a heterogeneous structure consisting of poly(ethylene glycol) dimethacrylate (PEG) hydrogels, which totally encase 100 μm diameter copper cylinders and 200 μm diameter polystyrene spheres. All the experiments were performed in a 20 mm × 20 mm × 4 mm chamber in phosphate-buffered saline (PBS). Snapshots of manipulation stages are shown in each subfigure, with the completed structure shown in schematic form in (g) and (l), corresponding to panes (e) and (k), respectively. Gels were placed on the second layer by moving them over a polyester plateau, which is the same thickness as the first layer of gels. The third layer was reached in (d) by pushing the gels up a polyester ramp. The time points of images are: 2:45 (a), 12:48 (b), 19:24 (c), 21:19 (d), 22:28 (e), 25:22 (f), 0:00 (h), 3:40 (i), 13:36 (j) and 15:39 (k) in minutes:seconds format. Scale bar, 1 mm.
And the assembly of shapes using mouse stem cells:
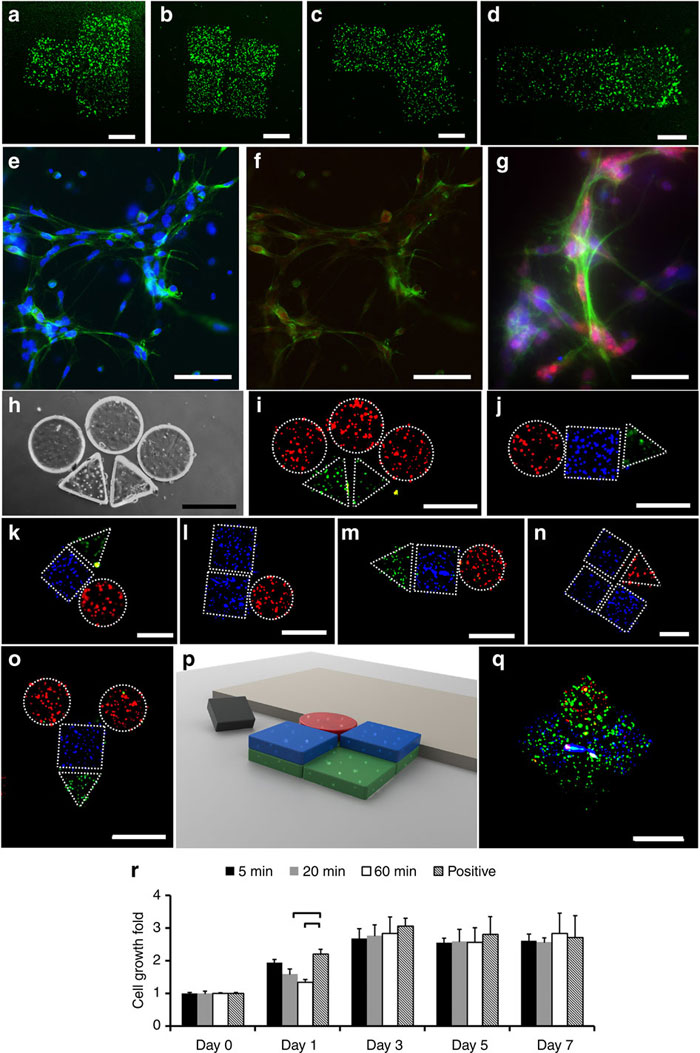
Fluorescence images of National Institutes of Health (NIH) 3T3 mouse embryonic fibroblast cell-encapsulating hydrogels after the assembly of (a) T-shape, (b) square-shape, (c) L-shape and (d) rod-shape constructs. Scale bar, 500 μm for (a–d). Green represents live cells and red represents dead cells. (e–g) Immunocytochemistry of proliferating cells stained with Ki67 (red), DAPI (blue) and Phalloidin (green) at day 4. (e) Cells stained with DAPI and Phalloidin at × 20 magnification. Scale bar, 100 μm. (f) Cells stained with Ki67 and Phalloidin at × 20 magnification. Scale bar, 100 μm. (g) Cells stained with Ki67, DAPI and Phalloidin at × 40 magnification. Scale bar, 40 μm. (a–g) Stainings were performed following the assembly of hydrogels. (h–q) Two- and three-dimensional heterogeneous assemblies of human umbilical vein endothelial cells, 3T3 and cardiomyocyte-encapsulating hydrogels. HUVECs, 3T3s and cardiomyocytes are stained with Alexa 488 (green), DAPI (blue) and propidium iodide (red), respectively. (h) Bright field and (i) fluorescence images of an assembly composed of circular and triangular gels. (j–o) Fluorescence images of several two-dimensional heterogeneous assemblies of HUVEC, 3T3 and cardiomyocyte-encapsulating hydrogels. (p) Schematic form and (q) fluorescence image of three-dimensional heterogeneous assembly of HUVEC, 3T3 and cardiomyocyte encapsulating hydrogels. Scale bar, 500 μm for (h–q). Stainings were performed before the assembly of hydrogels for (h–q). Teleoperated assembly durations of (a–d, h–q) are ~10 s to 5 min depending on the complexity of the final shape. (r) MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide, a yellow tetrazole) assay results of 3T3 cell suspensions in which micro-robots were kept for 5, 20 and 60 min durations. The positive control represents the cells that were incubated without any micro-robot presence. Results are normalized with day 0 absorbance values. Statistical analysis was only performed between positive control and (5, 20, 60) min cases. Brackets connecting groups indicate statistically significant difference (n=6, P<0.05). Error bars represent standard error of the mean.
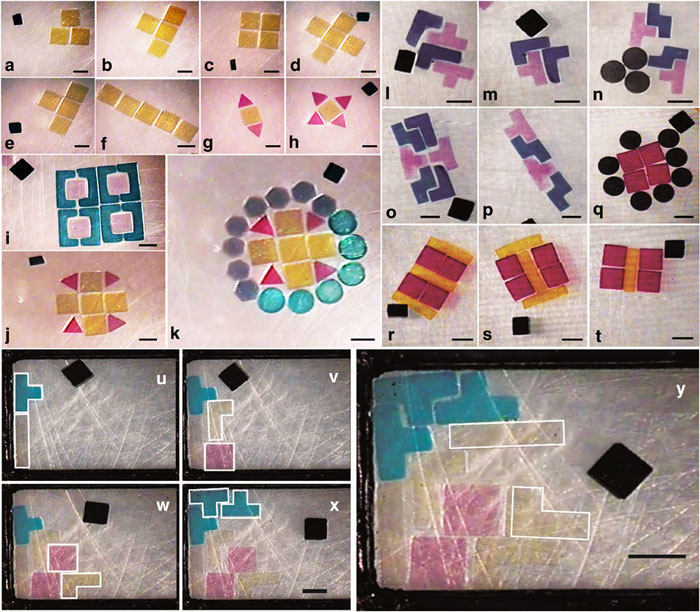
They also played Tetris just to show how precise they could be:
Micro-robotic coding and reconfiguration of Poly(ethylene glycol) dimethacrylate hydrogels (a–k) and gelatin methacrylate hydrogels (l–t) with various shapes into complex planar constructs. The black object in each image is top-view of a crawling micro-robot. To demonstrate the precision of micro-robotic manipulation, gels with several shapes including square, triangle, circle, hexagon, bracket-shape, plus-shape and others were coded. All the experiments were performed in 20 mm × 20 mm × 4 mm chamber in phosphate-buffered saline (PBS). Continuous coding and reconfiguring sequences are shown in panes (a–f), (g,h), (j,k), (l–p) and (r–t). Orientation and position control in untethered micro-robotic coding of material composition (u–y). Snapshots of ‘tetris’-shaped PEG hydrogels in a rectangular reservoir at different time points: 2:08 (u), 8:32 (v), 16:12 (w), 31:39 (x), 48:00 (y) in minutes:seconds format. Orientation and position of incoming hydrogels were dynamically changed as the geometry of cavities dynamically changed. All the experiments were performed in a 20 mm × 20 mm × 4 mm chamber. Scale bar, 1 mm.
Dr. Tasoglu explains that, “Compared with earlier techniques, this technology enables true control over bottom-up tissue engineering.” Their untethered microbot approach is still just the beginning of the group’s research, but CMU’s Metin Sitti hopes that their technique will change the field of 3D bioprinting, saying, “Our work will revolutionize three-dimensional precise assembly of complex and heterogeneous tissue engineering building blocks and serve to improve complexity and understanding of tissue engineering systems.”
Source: BWH